Bash/R basics
Contemporary molecular biology tasks are linked with big data generation, handling and analysis, making computational and programming skill development, essential. Within this context, students familiarize themselves with general concepts of essential tools for performing computational and biostatistics tasks, bash scripting and the R interpreter. Following, they are introduced to fundamental advanced sequencing and genomics concepts, both from the biology and the bioinformatics point of view, and gain hands-on experience with work examples.
Instructor: Sotirios Vasileiadis
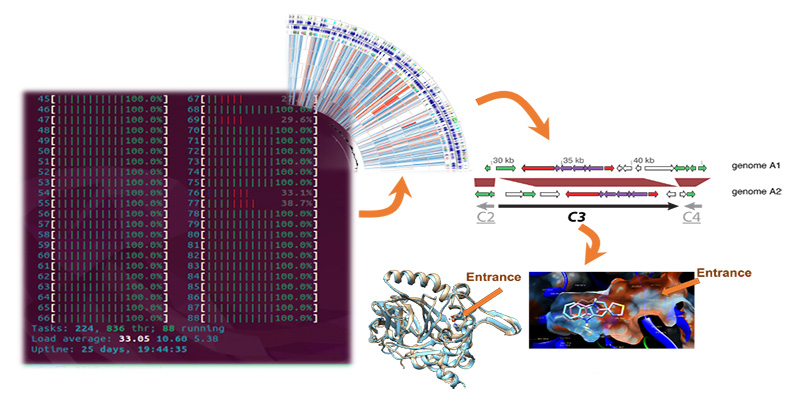
Genome assembly/annotation
Genomics comprises the basis of several systems biology analytical tasks. This module aims at providing all the necessary information for understanding automated genomic assembly in the era of high throughput sequencing, and how sequencing methods and reference genomes might affect it. Common strategies for computationally finishing small bacterial genomes and automated genome annotation approaches are also described. Students are expected to master this knowledge through worked examples and case studies.
Instructor: Sotirios Vasileiadis
RNAseq and ChIP sequencing
RNAseq and ChIP-seqare established and powerful methods of functional genomics, designed to interrogate the genome for gene expression and DNA-protein interactions, respectively. ChIP-seq results summarize the global binding profiles of transcription factors as well as the decoration of chromatin with epigenetic marks, in the form of histone modifications, ultimately leading to regulatory implications for the target loci. The aims of this module are: i) to mine publicly available ChIP-seq data (human and mouse genomes) and integrate them with genomic coordinates of regulatory importance (promoters, enhancers, etc) and/or RNA-seq results, ii) to functionally metanalyze them (both individually and comparatively) with the aim of distilling biological information and iii) to upload the results on genome browsers and to present them as high-quality images.
Instructor: Antonis Giakountis
Phylogenomics
In the rise of the post-genomic era, data analysis includes genome-wide phylogenetic markers to address evolutionary questions. During this module, students will learn how to disentangle phylogenetically noisy information and identify true orthologues, investigate paralogues of a gene family, and use this information for producing phylogenies out of genomic data.
Instructor: Alexie Papanicolaou
Single Cell genomics/transcriptomics
Finally, students take these topics at a different level, the level of single cells. During this module, students train on Single Cell Genomics, with an emphasis on technologies, methodologies, analytical tools, and project design. The instructor provides fundamental concepts along with case studies, next to a hands-on practice where students are required to answer in real-world datasets. The overall module aim is for the students to be able to identify cell subpopulations with distinct expression profiling characteristics and determine experimental treatment-induced changes in expression levels.
Instructor: Ioannis Ragoussis